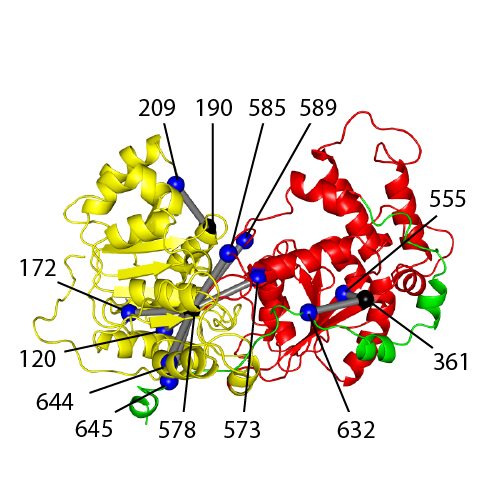
Probing the conformation of the ISWI ATPase domain with genetically encoded photoreactive crosslinkers and mass spectrometry
13-Dec-2011
Molecular & Cellular Proteomics, 2011, doi: 10.1074/mcp.M111.012088, published on 13.12.2011
Molecular & Cellular Proteomics, online article
Molecular & Cellular Proteomics, online article
We present a strategy for rapidly gaining structural information about a protein from crosslinks formed by genetically encoded unnatural amino acids. We applied it to ISWI, a chromatin remodeling enzyme involved in chromatin assembly, DNA replication and transcription. ISWI is part of the vast Snf2 family of helicase-related proteins, many of which constitute the catalytic cores of chromatin remodeling complexes. Structural information about this family is scarce, hampering our mechanistic understanding of chromatin remodeling. Making use of cells that harbour a special tRNA/aminoacyl-tRNA synthetase pair, several residues within the ATPase domain of ISWI were individually substituted with the UV-reactive unnatural amino acid p-benzoyl-p-phenylalanine. Intramolecular crosslinks could be mapped with amino acid precision by high resolution tandem mass spectrometry and the novel bioinformatic tool "Crossfinder". Most crosslinks were fully consistent with published crystal structures of ISWI-related ATPases. A subset of crosslinks, however, disagreed with the conformations previously captured in crystal structures. We built a structural model using the distance information obtained from the crosslinks and the structure of the closest crystallized relative, Chd1. The model shows the ATPase lobes strongly rotated against each other, a movement postulated earlier to be necessary to achieve a catalytically competent state. The minimal requirements for solubility and protein amounts make our approach ideal for studying structures and conformations of proteins that are not amenable to conventional structural techniques.